- Notebook: make changes to your module's code and see the effects immediately without needing to manually reload or restart the notebook's kernel.
%load_ext autoreload # enable the autoreload feature in your notebook session.
%autoreload 2 # sets the auto-reload mode to "2", which means that modules will be automatically re-loaded before executing any code that depends on them.- How to express
$10^{n}$ :2.944297e+10is equivalent to2.944297*(10**10)
- Modify the x-axis with highlighted values (only), instead of listing down all the dates
- In the below example, instead of putting in x-axis all the months, we just highlight data points with marks a year from 1949 to 1962
fig, ax = plt.subplots()
ax.plot(df['Month'], df['Passengers'])
ax.set_xlabel('Date')
ax.set_ylabel('Number of air passengers')
plt.xticks(np.arange(0, 145, 12), # only provide each 12 months
np.arange(1949, 1962, 1) # a list of a year corresponding to
)locgets rows (and/or columns) with particular labels.ilocgets rows (and/or columns) at integer locations.- Example: given the following dataframe that has the index starting from 80 to 84
df = pd.DataFrame(np.arange(25).reshape(5, 5),
index=[80, 81, 82, 83, 84],
columns=['col_A','col_B','col_C', 'col_D', 'col_E'])| col_A | col_B | col_C | col_D | col_E | |
|---|---|---|---|---|---|
| 80 | 0 | 1 | 2 | 3 | 4 |
| 81 | 5 | 6 | 7 | 8 | 9 |
| 82 | 10 | 11 | 12 | 13 | 14 |
| 83 | 15 | 16 | 17 | 18 | 19 |
| 84 | 20 | 21 | 22 | 23 | 24 |
- Return the first row of the df with two columns
col_Aandcol_D.loc: since the first row in the dataframe corresponding to theindex=80, so we need to specify it in the.locdf.loc[80, ["col_A", "col_D"]]
.iloc: as iloc will be based on the integer location, so the first row in the df is corresponding to the location 0df.iloc[0, [df.columns.get_loc(c) for c in ["col_A", "col_D"]]]
# example of .loc and .iloc to return the first row of the df with two columns A and D
df.loc[80, ["col_A", "col_D"]]
# col_A 0
# col_D 3
df.iloc[0, [df.columns.get_loc(c) for c in ["col_A", "col_D"]]]
# col_A 0
# col_D 3- Select last 2 row in the
df→.ilocwill have the advantages as it is based on the integer location rather then the labels.
df.iloc[-2:, :]| col_A | col_B | col_C | col_D | col_E | |
|---|---|---|---|---|---|
| 83 | 15 | 16 | 17 | 18 | 19 |
| 84 | 20 | 21 | 22 | 23 | 24 |
- Select first 3 columns of the row after index=82
df.iloc[df.index.get_loc(82):, :3]| col_A | col_B | col_C | |
|---|---|---|---|
| 82 | 10 | 11 | 12 |
| 83 | 15 | 16 | 17 |
| 84 | 20 | 21 | 22 |
1e-4is equal to0.0001with total 4 zeros- Validate the input string variable, say
splitbelongs to certain options:assert split in ['train', 'test', 'both']- We can add the handling path if the assertion error paused the program
try: assert 'value' not in numerical_vars except AssertionError: # do something
- The
meltfunction in pandas is used to unpivot a DataFrame, meaning that it converts columns of data into rows.- This is useful when you want to combine the data for plotting
- For example: the melt function has converted the
MathandPhysicscolumns into rows, and created two new columns:subjectandscore.
# Create a sample DataFrame
df = pd.DataFrame({'student_id': [1, 2, 3], 'Math': [4, 5, 6], 'Physics': [7, 8, 9]})
# Melt the DataFrame
df_melted = df.melt(id_vars=['student_id'], value_vars=['Math', 'Physics'], var_name='Ssubject', value_name='score')
# Print the melted DataFrame
print(df_melted)
# student_id subject score
# 0 1 Math 4
# 1 2 Math 5
# 2 3 Math 6
# 3 1 Physics 7
# 4 2 Physics 8
# 5 3 Physics 9
# plotting the melted dataframe by subject using hue of seaborn
sns.barplot(df_melted, x='student_id', y='score', hue='subject')- The
crosstabfunction: produces a DataFrame where the rows represent the levels of one factor and the columns represent the levels of another factor. - Usage: to create frequency table for each category in a feature vs another feature
- The crosstab function takes the following arguments:
index: The name of the column to use for the row labels.columns: The name of the column to use for the column labels.values: The name of the column to use for the cell values. If not specified, the counts of observations are used.margins: If True, the DataFrame will include a row and column for the totals.
# Create a sample DataFrame
df = pd.DataFrame({'gender': ['M', 'F', 'M', 'F', 'F'], 'favorite_color': ['blue', 'red', 'green', 'blue', 'purple']})
# gender favorite_color
# 0 M blue
# 1 F red
# 2 M green
# 3 F blue
# 4 F purple
# Create a crosstabulation of gender and favorite color
crosstab = pd.crosstab(df['gender'], # rows
df['favorite_color'], # columns
margins=True # True to calculate the total
)
# re-name columns "All" to 'row_totals' & 'col_totals'
# note: columns "All" are only avail if margins=True
crosstab.columns = [*crosstab.columns.to_list()[:-1], 'row_totals']
crosstab.index = [*crosstab.index.to_list()[:-1], 'col_totals']
# blue green purple red row_totals
# F 1 0 1 1 3
# M 1 1 0 0 2
# col_totals 2 1 1 1 5- Subplots:
# Annual, weekly and daily seasonality
# ==============================================================================
fig, axs = plt.subplots(2, 2, figsize=(8.5, 5.5), sharex=False, sharey=True)
# Before:
ax1 = axs[0,0]
ax2 = axs[0,1]
# After with axs.ravel()
axs = axs.ravel()
ax1 = axs[0]
#...
ax4 = axs[3]- Pip installs packages from the Python Package Index (
PyPI), which hosts a vast array of Python libraries. Almost any Python library can be installed using pip. - conda installs packages from the Anaconda distribution and other channels (
conda-forge). While the number of packages available through conda is smaller than pip, conda can install packages for multiple languages and not just Python. - Example:
skforecastis not avail in theconda-forgebut it is avail inPyPI, so you cannotconda install skforecast, but it can be install viapip installcommand
- Find the row with the column is equal to value:
index_choice = df.index.get_loc(CustomerId=15674932)
| Space between 2 bins | Frequency of Samples in each bin | |
|---|---|---|
pd.cut |
Equal Spacing | Difference |
pd.qcut |
Un-equal Spacing | Same |
pd.cutwill choose the bins to be evenly spaced according to the values themselves and not the frequency of those values.- You also can define the bounds for each binning with
pd.cut() - You can use the Fisher-Jenks algorithm to determine the natural bounds and then pass those values into
pd.cut()
- You also can define the bounds for each binning with
pd.qcutthe bin interval will be chosen so based on the percentiles that you have the same number of records in each bin.
factors = np.random.randn(30)
pd.cut(factors, 5).value_counts() # bin size has equal interval of ~1
# (-2.583, -1.539] 5
# (-1.539, -0.5] 5
# (-0.5, 0.539] 9
# (0.539, 1.578] 9
# (1.578, 2.617] 2
pd.qcut(factors, 5).value_counts() # each bin has equal size of 6
# [-2.578, -0.829] 6
# (-0.829, -0.36] 6
# (-0.36, 0.366] 6
# (0.366, 0.868] 6
# (0.868, 2.617] 6
- If the csv file is large, can consider to read by chunk
df_iter = pd.read_csv(file_path, iterator=True, chunksize=100000)
df = next(df_iter)@lru_cachemodifies the function it decorates to return the same value that was returned the first time, instead of computing it again, executing the code of the function every time.
@lru_cache
def say_hi(name: str, salutation: str = "Ms."):
return f"Hello {salutation} {name}"Annotatedin python allows developers to declare the type of a reference and provide additional information related to it.
from typing_extensions import Annotated
# This tells that "name" is of type "str" and that "name[0]" is a capital letter.
name = Annotated[str, "first letter is capital"]- Fast API examples:
from fastapi import Query def read_items(q: Annotated[str, Query(max_length=50)])
- The parameter
qis of typestrwith a maximum length of 50.
- The parameter
- Convert
floatdirectly toDecimalconstructor introduces a rounding error.
from decimal import Decimal
x = 0.1234
Decimal(x)
# Decimal('0.12339999999999999580335696691690827719867229461669921875')- Solution: to convert a float to a string before passing it to the constructor.
- You also can round the float before converting it to string
Decimal(str(x))
# Decimal('0.1234')
Decimal(str(round(x,2)))
# Decimal('0.12')-
subprocess.runis a higher-level wrapper around Popen that is intended to be more convenient to use.- Usage: to run a command and capture its output
-
subprocess.call- Usage: to run a command and check the return code, but do not need to capture the output.
-
subprocess.Popenis a lower-level interface to running subprocesses- Usage: if you need more control over the process, such as interacting with its input and output streams.
-
A
PIPEis a unidirectional communication channel that connects one process's standard output to another's standard input.
# creates a pipe that connects the output of the `ls` command to the input of the `grep` command,
ls_process = subprocess.Popen(["ls"], stdout=subprocess.PIPE, text=True)
grep_process = subprocess.Popen(["grep", ".ipynb"], stdin=ls_process.stdout, stdout=subprocess.PIPE, text=True)
output, error = grep_process.communicate()
print(f"Ouptut:\n{output}")
print(f"Error : {error}")
# Ouptut:
# google_colab_tutorial.ipynb
# subprocess.ipynb
#
# Error : None
result = subprocess.run(["ls"], stdout=subprocess.PIPE)
print(result.stdout.decode()) # decode() to convert from bytes to strings
# google_colab_tutorial.ipynb
# subprocess.ipynbsubprocess.runis generally more flexible thanos.system(you can get the stdout, stderr, the "real" status code, better error handling, etc.)- Even the documentation for
os.systemrecommends using subprocess instead.
- The kernel knows to execute this script with a python interpreter with the shebang
#!/usr/bin/env python sys.argv[0]return name of the scriptsys.argv[1:]return the arguments parsed to the script
#############################
# in the python_script.py #
#############################
#!/usr/bin/env python
import sys
for arg in reversed(sys.argv[1:]):
print(arg)
#############################
# in the interactive shell #
#############################
bash-5.2$ chmod +x python_script.py
bash-5.2$ ./python_script.py a b c
# Running: ./python_script.py
# c
# b
# abytes("yes", "utf-8")convert string to binary objects:
- Color Map
cmap = plt.get_cmap("viridis")
fig = plt.figure(figsize=(8, 6))
m1 = plt.scatter(X_train, y_train, color=cmap(0.9), s=10)
m2 = plt.scatter(X_test, y_test, color=cmap(0.5), s=10)- Plot 2 charts on the same figure with share x-axis
fig, ax1 = plt.subplots()
ax1.hist(housing["housing_median_age"], bins=50)
ax2 = ax1.twinx() # key: create a twin axis that shares the same x-axis
color = "blue"
ax2.plot(ages, rbf1, color=color, label="gamma = 0.10")
ax2.tick_params(axis='y', labelcolor=color)
ax2.set_ylabel("Age similarity", color=color) # second y-axis's measurement
plt.show()# example.py
from .abstract import ExampleClass
# if we run this script directly like python src/abc/example.py, we will encounter the issue
- Soluton:
python -m src.abc.exampleor call theexample.pyscript outside thesrc
from subprocess import Popen, Pipe
execute = Popen("scp -i /location_in_machine_A/to/private/key -v -P 64022 file_path machineB@ip_address:/path/to/store/inB/".split(), stdout=PIPE, stdin=PIPE, stderr=PIPE)
execute.stdin.write(bytes("yes", "utf-8")) # to send the "yes" command
execute.communicate()[0]- To read the path separator of the env
os.path.sep # return '/' if using Linux
- Copy a list:
copied_list = a_list[:]- Leverage on
copymodule
import copy copied_list = copy.deepcopy(a_list)
- Leverage on
- Conda is an open source package + environment manager
- Conda vs (Miniconda & Anaconda):
- Conda is a package manager & Conda is tightly coupled to two software distributions: Anaconda and Miniconda.
- Anaconda is a full distribution of the central software in the PyData ecosystem, and includes Python itself along with binaries for several hundred third-party open-source projects.
- Miniconda is essentially an installer for an empty conda environment, containing only Conda and its dependencies, so that you can install what you need from scratch.
- Recommended conda installation in PC: Miniconda
- Miniconda and Miniforge:
- Miniforge-installed conda is the same as Miniconda-installed conda, except that it uses the conda-forge channel (and only the conda-forge channel) as the default channel.
- Miniconda-installed conda is the Anaconda (company) driven minimalistic conda installer, where pacakages installed from the anaconda channels
- Conda channel: A channel is the location where packages are stored remotely.
- Example of channels: Anaconda channel,
conda-forge, Apple - Install TensorFlow dependencies from Apple Conda channel:
conda install -c apple tensorflow-deps
- Example of channels: Anaconda channel,
- Understanding
environment.ymlinconda create --prefix ./venv python=3.8 --file environment.ymltensorflow-depswill need to be installed bycondaviaapplechannelscikit-learnwill be installed bycondaviaconda-forgechanneltensorflow-macoswill needpipinstall via PyPI
name: tensorflow
channels:
- apple
- conda-forge
dependencies:
- tensorflow-deps
- scikit-learn
- pip:
- tensorflow-macos
- tensorflow-metal# create env
conda create --prefix ./venv python=3.8 --file requirements.txt
conda activate ./venv
conda deactivate
# install package from requirements.txt
conda install --file requirements.txt
# export to requirements.txt so that can install via pip in virtualenv/venv environment
pip list --format=freeze > requirements.txtconda create --fileexpects arequirements.txt, not anenvironment.yml, each line in the given file is treated as a package-referenceconda env createto create an environment from a givenenvironment.yml- Command:
conda env create --name tensorflow --file environment.yml
- Command:
conda env listto list down conda envsdu -h -s $(conda info --base)/envs/*list down the size of each envconda config --show channelsto show available channels in the config
- Number:
1000000can be written as1_000_000for the ease of visualisationstate['Population'] / 1_000_000
- Plot horizontal line:
{plt, ax}.axhline(y=0.5, color='r', linestyle='-')
- Stacking columns/rows
np.column_stack&np.row_stacknp.hstack&np.vstack
a = np.array((1,2,3))
b = np.array((2,3,4))
# column stack
np.column_stack((a,b))
# array([[1, 2],
# [2, 3],
# [3, 4]])
# row_stack or vstack
np.row_stack((a,b))
np.vstack((a,b))
# array([[1, 2, 3],
# [2, 3, 4]])
# hstack
np.hstack((a,b))
# array([1, 2, 3, 2, 3, 4])- Pandas's holiday package:
pandas.tseries.holiday holidayspackage
- Instead of
if '.yml' in file_path or '.yaml' in file_pathwe can do as follows:- Solution:
if any(ext in file_path for ext in ['.yml', '.yaml']
- Solution:
- To get color palatte
color_pal = sns.color_palette()
- Since the
sns.displotis deprecated, so we will usesns.histplotas follow to plot the histogram + kde distribution by specifyingkde=True - Also can
set_xlim()to the zone that containing the data in case the distribution is skewed
sns.histplot(df['col_name'], kde=True, bins=50).set_xlim(0,8);- Pairplot is to use
scatterplot()for each pairing of the variables andhistplot()for the marginal plots along the diagonal - Customise with
x_varandy_varandhue
sns.pairplot(df.dropna(),
hue='hour',
x_vars=['hour','dayofweek',
'year','weekofyear'],
y_vars='PJME_MW',
height=5,
plot_kws={'alpha':0.15, 'linewidth':1.5}
)
plt.suptitle('Power Use MW by Hour, Day of Week, Year and Week of Year')
plt.show()yieldkeyword is used in the context of defining generator functions.- When a generator function is called, it doesn't execute the function immediately.
- Instead, it returns a generator object that can be used to control the execution of the function.
- The code inside the generator function is executed only when an item is requested from the generator using the
next()function or aforloop.
def simple_generator():
yield 1
yield 2
yield 3
# Create a generator object
gen = simple_generator()
# using next() to access the code inside
print(next(gen)) # 1
print(next(gen)) # 2
print(next(gen)) # 3
print(next(gen)) # StopIteration raised
# using for-loop to iterate through the generator
for value in simple_generator():
print(value)- Numerical columns:
num_cols = df.select_dtypes(include=np.number).columns.tolist() - Categorical columns:
cat_cols = df.select_dtypes(exclude=np.number).columns.tolist()
- Convert the datetime index to a datetime col:
df['date'] = df.index.date - Slicing using 1 date by including
23:59:59
end_train = '1980-01-01 23:59:59'
# including 23:59:59 means
data_train = df.loc[:end_train] # ends at 1980-01-01
data_test = df.loc[end_train:] # start at 1980-01-02- Can set multiple condition:
df.query("make == 'bmw' and model == '1_series') - Can query using a list
holiday_list = ['2021-01-01', '2022-09-02']
df.query('datetime_col in @holiday_list')df.duplicated()to check if there are any row duplicate. This will returnTruefor the 2nd occurence of the duplicatedf.duplicated(subset=['col_A','col_B','col_C'])in case there is no entire row duplciate, we can check duplicates for only subsets of columns
df.query("make == 'bmw' and model == '1_series' and year == 2013 and price == 31500")using query to identify & view the duplicated rows- Remove the duplicates
.reset_index(drop=True)to reset the index after dropping the duplicates.copy()to make the deep copy of the dataframe
df = df.loc[~df.duplicated(subset=['Coaster_Name','Location','Opening_Date'])] \
.reset_index(drop=True).copy()- You can set matplotlib object to
axvariable- You also can continue to plot on the same graph with
axvariable
- You also can continue to plot on the same graph with
# case 1: get ax from the plot via pandas dataframe
ax = df['year'].value_counts() \
.head(10) \
.plot(kind='bar', title='Top 10 Years Coasters Introduced')
# case 1.1: also can continue to plot on the same graph with ax variable
df.query('year < 2023').plot(style='.', ax=ax)
ax.set_xlabel('year')
ax.set_ylabel('count')
# case 2: get ax from the plot via seaborn
ax = sns.countplot(data=df, x='year')
ax.set_xticks(ax.get_xticks(), ax.get_xticklabels(), rotation=90, ha='center')- Rotate the xticks label
# rotate via plt
plt.xticks(rotation=90)
# rotate via ax
ax.set_xticks(ax.get_xticks(), ax.get_xticklabels(), rotation=45, ha='right')- Numpy 's
np.nanvs Python 'sNoneobject- In Numpy, a
np.nanvalue is a native floating-point type array.- When you try to do some arithmetic operations will
np.nan, the result will always benp.nan. - Fortunately, Numpy provides some special aggregation methods that can ignore the existence of
np.nanvalue such asnp.nansum(arr)
- When you try to do some arithmetic operations will
Noneis a Python Object called NoneType- Pandas automatically converts the
Noneto anp.nanvalue. - If you try to aggregate over this array, you will get an error because of the NoneType.
- Pandas automatically converts the
- In Numpy, a
df.loc[:, "col"] = df["col"].map(mapping)re-assign the updated value to originial column without any errorpd.options.display.max_columnsto view all the columns in the df whendf.head()
- Use the
.aggfunction to get multiple statistics on the other columns
## Example 1:
df.groupby('A').agg(['min', 'max']) # apply same operations on other columns
## Example 2:
df.groupby('A').agg({'B': ['min', 'max'], 'C': 'sum'}) # apply different operations on other columns
## Example 3:
# group by Team & Position, get mean, min, and max value of Age for each value of Team.
grouped_single = df.groupby(['Team', 'Position']).agg({'Age': ['mean', 'min', 'max']})
grouped_single.columns = ['age_mean', 'age_min', 'age_max'] # rename columns
# reset index to get grouped columns back
grouped_single = grouped_single.reset_index()
## Example 4:
df_customers.groupby('rfm_score').agg( # groupby 'rfm_score' col
customers=('customer_id', 'count'), # create new 'customers' col by count('customer_id' col)
mean_recency=('recency', 'mean'), # create new 'mean_recency' col by mean('recency' col)
mean_frequency=('frequency', 'mean'),
mean_monetary=('monetary', 'mean'),
).sort_values(by='rfm_score')- Group by the first column and get second column as lists in rows using
.apply(list)
In [1]: df = pd.DataFrame( {'a':['A','A','B','B','B','C'], 'b':[1,2,5,5,4,6]})
df
Out[1]:
a b
0 A 1
1 A 2
2 B 5
3 B 5
4 B 4
5 C 6
In [2]: df.groupby('a')['b'].apply(list)
Out[2]:
a
A [1, 2]
B [5, 5, 4]
C [6]
Name: b, dtype: object-
IPythondebug: when executingmain.pyscript in the terminal, we still can insert ipython checkpoint at the line we want to debugfrom IPython import embed; embed()
-
Avoid circular imports
# __init__.py of utils folder from .base_logger import * from .data_loader import * def load_yaml() # data_loader.py from utils import load_yaml # this will cause circular import- Solution: DO NOT
from .data_loader import *if a data_loader refer to any functions inutils.__init__.py
- Solution: DO NOT
- Installation:
pip install python-dotenv==1.0.0 - Config file stores in
.envfile
HUGGINGFACEHUB_API_TOKEN="<hf_token>"- Load environmental variables
import os
from dotenv import load_dotenv, find_dotenv
load_dotenv("../config/.env")
# load_dotenv(find_dotenv()) # find_dotenv() is to find the .env
os.environ["HUGGINGFACEHUB_API_TOKEN"] = ... # insert your API_TOKEN here- Surpass the warning
import warnings
warnings.filterwarnings('ignore')
# [Optional] If you do not want to supress all the warnings, you also can explicitly specify which warning needs ignore
import warnings
warnings.filterwarnings('ignore', category=FutureWarning)
warnings.filterwarnings('ignore', category=DeprecationWarning)- Both
!and%allow you to run shell commands from a Jupyter notebook- Difference:
!calls out to a shell (in a new process), while%affects the process associated with the notebook!cd foo, by itself, has no lasting effect, since the process with the changed directory immediately terminates.%cd foochanges the current directory of the notebook process, which is a lasting effect.
- Example
# in a notebook cell !git clone https://github.com/full-stack-deep-learning/fsdl-text-recognizer-2021-labs # ! can use for git clone, pip install %cd fsdl-text-recognizer-2021-labs # % use for cd
- Difference:
- Multiline editing in Visual Studio Code:
- Mac:
⌥ Opt+⌘ Cmd+↑/↓ - Windows:
Shift+Alt+↑/↓
- Mac:
- Figure object is the overall window where everything is drawn.
def draw_smtg():
fig = plt.figure(figsize=(10,10))
plt.plot(...)
return fig #whatever the graphs in fig will be returned- Logging level:
DEBUG > INFO > WARNING > ERROR > CRITICAL- If set
logging.basicConfig(level=logging.ERROR)means that only logERROR&CRITICAL
- If set
reloada module in Jupyter notebook
from importlib import reload
from abc import module_a
# By doing this, whatever code change in class_xyz will be reflected in the notebook
reload(module_a)
from abc.module_a import class_xyzpython -m pip install <some-package>-mflag makes sure that you are using the pip that's tied to the active Python executable.
- Load pickle file using
joblib
model = joblib.load('lgbm_mode.pkl')df = pd.read_csv('example.csv',index_col=[0], parse_dates=[0])to set the col loc=0 as the index, and parsed as date time typeto_csvto preventnnamed: 0column to be appended along with your original df by setdf.to_csv('result.csv', index=False).applybased on the condition of certain columns
df.loc[:,'C'] = df.apply(lambda row: 'Hi' if row['A'] > 10 and row['B'] < 5 else '')df.insert()Avoid errorTry using .loc[row_indexer,col_indexer]when creating the new column in df from an existing df
# insert(position of the newly_inserted_col in the df, newly_inserted_col's name, newly_inserted_col's value)
data.insert(len(data.columns), 'rolling', data['open'].rolling(5).mean().values)- Concat
dfwithnumpy_arrayand assign the name for thenumpy_arrayasnew_col- Syntax:
df = df.assign(new_col=numpy_array)
- Syntax:
# need to convert pd's Series into the Dataframe, and transpose it before concat with pd's DF
pd.concat([df, pd_series.to_frame().T], ignore_index=True)- Experience: before joining (either
concat,merge), need to careful about the index of the dataframes (might need to.reset_index()) as apart from the joining condition, pandas also matching the index of each dataframe.
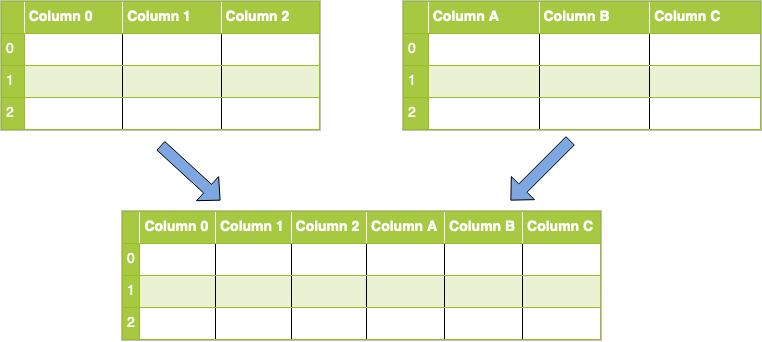
Column Concat
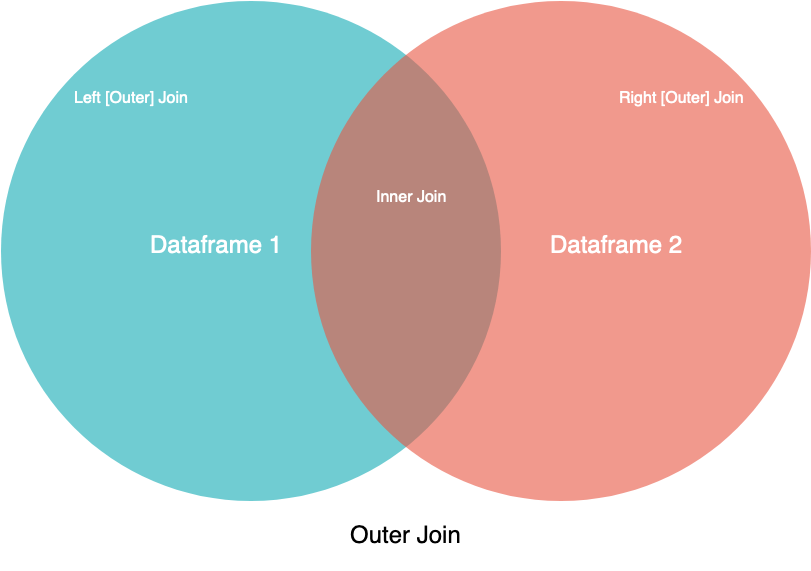
Merge [Inner, Outter (Left, Right)]
concat()for combining DataFrames across rows or columns- axis represents the axis that you’ll concatenate along.
- The default value is 0, which concatenates along the index, or row axis.
- Alternatively, a value of 1 will concatenate vertically, along columns. You can also use the string values "index" or "columns".
ignore_indexdefaults to False.- If True, then the new combined dataset won’t preserve the original index values in the axis specified in the axis parameter. This lets you have entirely new index values.
- axis represents the axis that you’ll concatenate along.
merge()for combining data on common columns or indices- how defaults to
inner, but other possible options includeouter,left, andright on;left_onandright_onspecify a column or index that’s present only in the left or right object that you’re merging
- how defaults to
# ------------ pd.concat() examples ------------
reindexed = pd.concat(
[df1, df2], ignore_index=True, axis=1
)
# ------------ pd.merge() examples -------------
pd.merge(
df1, df2, how="left", on=["col_A", "col_B"]
)
# if left & right DF has diff joining col names
pd.merge(
df1, df2, how="left", left_on=["col_A1"], right_on=["col_A2"]
)- Dense & Sparse Matrix Conversion:
- Sparse → Dense:
A.toarray()
- Sparse → Dense:
- Numpy vs Tensor:
- TensorFlow seems to look a lot like NumPy. But here’s something NumPy can’t do: retrieve the gradient of any differentiable expression with respect to any of its inputs.
- Open a
GradientTapescope, apply some computation to one or several input tensors, and retrieve the gradient of the result with respect to the inputs.
- Open a
- A significant difference between NumPy arrays and TensorFlow tensors is that TensorFlow tensors aren’t assignable: they’re constant
- TensorFlow seems to look a lot like NumPy. But here’s something NumPy can’t do: retrieve the gradient of any differentiable expression with respect to any of its inputs.
import numpy as np
x = np.ones(shape=(2, 2))
x[0, 0] = 0.
x = tf.ones(shape=(2, 2))
x[0, 0] = 0. # ERROR: fail, as a tensor isn’t assignable.sys.pathis a built-in variable within the sys module. It contains a list of directories that the interpreter will search in for the required module. When a module(a module is a python file) is imported within a Python file, the interpreter first searches for the specified module among its built-in modules. If not found it looks through the list of directories(a directory is a folder that contains related modules) defined by sys.path.- To locate the installation path of the module:
print(module_name.__file__) - Python will locate the module based on the path appears first in
sys.path, so in order to change prioritise the installation packages, we can do as follow:
import sys
sys.path.insert(0, '/path/to/site-packages') # location of src- Set params:
plt.rcParams.update({'font.size': 14}) - Set the style of the plot
plt.style.use('fivethirtyeight') # set at the front of the notebook- Other styles:
seaborn-v0_8-darkgrid
- Other styles:
- Set vertical axis range:
ax.set_ylim([0,1])orplt.gca().set_ylim(0,1) #set vertical range to [0-1] - Set horizontal axis range:
- Enable subplots share the same axis with
sharexorsharey:plt.subplots(nrows=3, sharey=True) fig.tight_layout(pad=2)function of matplotlib allows to adhust the gap between subplotspadparameter to specify gap size
assert: to check if the data is expected (assert len(x.shape) == 2) and will raise Exception if not matching.iter(): return an iterator for the given a list, set, tuple or object with__next__()method.
phones = ['apple', 'samsung', 'oneplus']
phones_iter = iter(phones)
print(next(phones_iter))os.environ[variable]=valuea mapping object that represents the user’s environmental variables.sys.pathis a built-in variable within the sys module. It contains a list of directories that the interpreter will search in for the required module. When a module(a module is a python file) is imported within a Python file, the interpreter first searches for the specified module among its built-in modules. If not found it looks through the list of directories(a directory is a folder that contains related modules) defined by sys.path.randommodule- Randomly select an item in a list:
random.randint(0, len_x) - Randomly select subset of items in a list:
# Method 1 indices_permutation = random.permutation(len(x)) dataset[indices_permutation][:10] # Method 2: select 10 out of the dataset indices_permutation = random.sample(range(len(trainset)), 10) dataset[indices_permutation]
- Normal distribution with mean 0 and standard deviation 1:
np.random.normal(size=(3,1), loc=0., scale=1.) - Uniform distribution between 0 and 1:
np.random.uniform(size=(3, 1), low=0., high=1.)
- Randomly select an item in a list:
CMD + Shift + PCommand PaletteCMD + PQuickly open files
# in .vscode settings.json
"python.terminal.activateEnvironment": true- The main coding standard for Python is PEP8 (Python Enhancement Proposal 8)
- Linters such as
flake8andpylinthighlight places where your code doesn’t conform with PEP8. - Automatic formatters such as
blackthat will update your code automatically to conform with coding standards. - Type Checker
mypyis a static type checker for Python. Type checkers help ensure that you're using variables and functions in your code correctly. With mypy, add type hints (PEP 484) to your Python programs, and mypy will warn you when you use those types incorrectly.
- Linters such as
- How to install
flack8:- Step 1: Install
blackin virtual environment:pip install flake8 - Step 2: Open the Command Palette (
CMD + Shift + P) → Search the “Python: Select Linter” and press enter. Select the “flake8”
- Step 1: Install
- How to install
black:- Step 1: Install
blackin virtual environment:pip install black - Step 2: Open the Command Palette (
CMD + Shift + P) → “Preferences → Settings”- Search “format on save” and check the checkbox
- Search “python formatting provider” and select the
black.
- NOTE: can run black seperately
black -l 80 --preview src/youtube_statistics.py
- Step 1: Install
- How to install
isort:- Open the Command Palette (
CMD + Shift + P). Search the “Preferences: Open User Settings (JSON)” and press enter. It will open the “settings.json” and add the follow code into that:
"editor.codeActionsOnSave": { "source.organizeImports": true }
- Open the Command Palette (
This is to ensure the code formatter, linting is running before the commit
- install requirements-dev dependencies
# requirements-dev.txt
flake8
black
mypy
coverage-
add
hooksPathinto git config core:git config --local core.hooksPath .git/hooks/ -
pre-commitfile inside.git/hooks/# content inside pre-commit file echo "Running lint.sh before commit" bash lint.sh echo "Linting completed"
- make pre-commit file executable via
chmod +x .git/hooks//pre-commit
- make pre-commit file executable via
-
Define the content in
lint.sh# content inside lint.sh linting_path="." echo "-------------Formatter with black-----------------" black $linting_path --line-length=88 echo "-------------Linting with flake8-----------------" flake8 $linting_path --max-line-length=88 --ignore=E203,W503,E231,E266,E722 echo "-------------Type checking with mypy-----------------" mypy $linting_path --ignore-missing-imports

