This is the contribution of the CDS team to the MedGemma Impact Challenge 2026.
Team: Clinical Data Science in Radiology, Department of Radiology, LMU University Hospital, LMU Munich License: CC BY 4.0
- Get an Openai compatible URL and API key
- Configure and start:
Edit code/agent-api/.env — paste your API key as LLM_API_KEY and EMBEDDING_API_KEY and your URL as LLM_BASE_URL and LLM_EMBEDDING_URL as well as your embedding model.
cp code/agent-api/.env.example code/agent-api/.env
cd code docker compose build && docker compose up
- Open the app at http://localhost:3000 after the application startup has completed.
At the Clinical Data Science Group in the Department of Radiology at LMU Hospital Munich, we have built what our clinicians had long been wishing for: a more innovative, faster way to prepare and run tumor boards. In a few intense hackathon weeks, together with practicing physicians and hospital IT engineers, we developed TURBO. This MedGemma-based agentic tumor board assistant mirrors real-world workflows and aligns with institutional governance requirements.
- Clinical Data Science, Radiology: Prof. Dr. Michael Ingrisch, Jakob Dexl, David Götzinger, Dr. Katharina Jeblick, Anna Theresa Stüber, Johanna Topalis, Rachelle Trotman, Beatrice Villata
- Hospital IT: Gabriel Lindner, Stefan Reifberger, Dr. Balthasar Schachtner
- Clinical advisors: Prof. Dr. Bastian Sabel (Senior Radiologist), Prof. Dr. Amanda Tufman (Senior Specialist in Thoracic Oncology)
Multidisciplinary tumor boards are a central decision-making structure in oncology. For each case, clinicians must reconstruct a longitudinal narrative from radiology reports, pathology, prior therapies, comorbidities, and current oncological guideline recommendations to define the best possible treatment plan. Although these data are available digitally, they are fragmented across heterogeneous hospital systems and are stored mainly as unstructured text. Preparation is therefore manual, time-intensive, and cognitively demanding. Physicians retrieve documents from multiple sources, assess their clinical relevance, integrate findings across time, and synthesize a coherent summary under time pressure. At German university hospitals, tumor board preparation and participation can exceed 142 min, including 53 min preparation per tumor board per week [1]. This manual workflow has two consequences: reduced time for patient-facing care and an increased risk that relevant longitudinal details will be overlooked in complex cases. As oncology becomes more data-rich, this burden will intensify.
Tumor board preparation is well-suited for domain-specific foundation models. It requires not only cross-document synthesis, contextual relevance assessment, and structured summarization but also medically grounded reasoning across all patient data and oncological guidelines. A medically pre-trained, on-premise deployable model can systematically aggregate and contextualize patient data, surface clinically meaningful connections, and support hypothesis-driven discussion, reducing manual preparation time while maintaining human oversight.
We are not alone in seeing this potential. A major pharmaceutical company has invested heavily in digitizing tumor board workflows, most recently testing LLM-based summarization on its commercial platform [2]. Potential time and financial gains are substantial. Our senior physician estimated a 50% reduction in preparation time, consistent with the company's claims of a 40–50% reduction [3]. For a per-case preparation time between 4 min [1] and 30 min [3], a 50% reduction translates to €36K–€270K per university hospital, assuming €100/hr physician cost (16.7 boards/week × 13.5 cases/board × 48 weeks = ~10,800 cases/yr [1]). With roughly 20 million new cancer cases annually [4] and tumor boards held in most high-income countries [5], we conservatively estimate 10M tumor board preparations per year worldwide, translating to €40M–€300M globally. These numbers are expected to rise significantly, as cancer incidence is projected to reach 35 million cases annually by 2050 [4].
This is why we have developed TURBO, an agentic, on-premise tumor board assistant built around MedGemma as its clinical reasoning engine. While maintaining complete human oversight, the system streamlines tumor board case preparation and discussion by transforming fragmented patient records into structured, verifiable, and discussion-ready case summaries and enabling case-specific follow-up questions via an integrated agentic chatbot.
The solution comprises three integrated components:
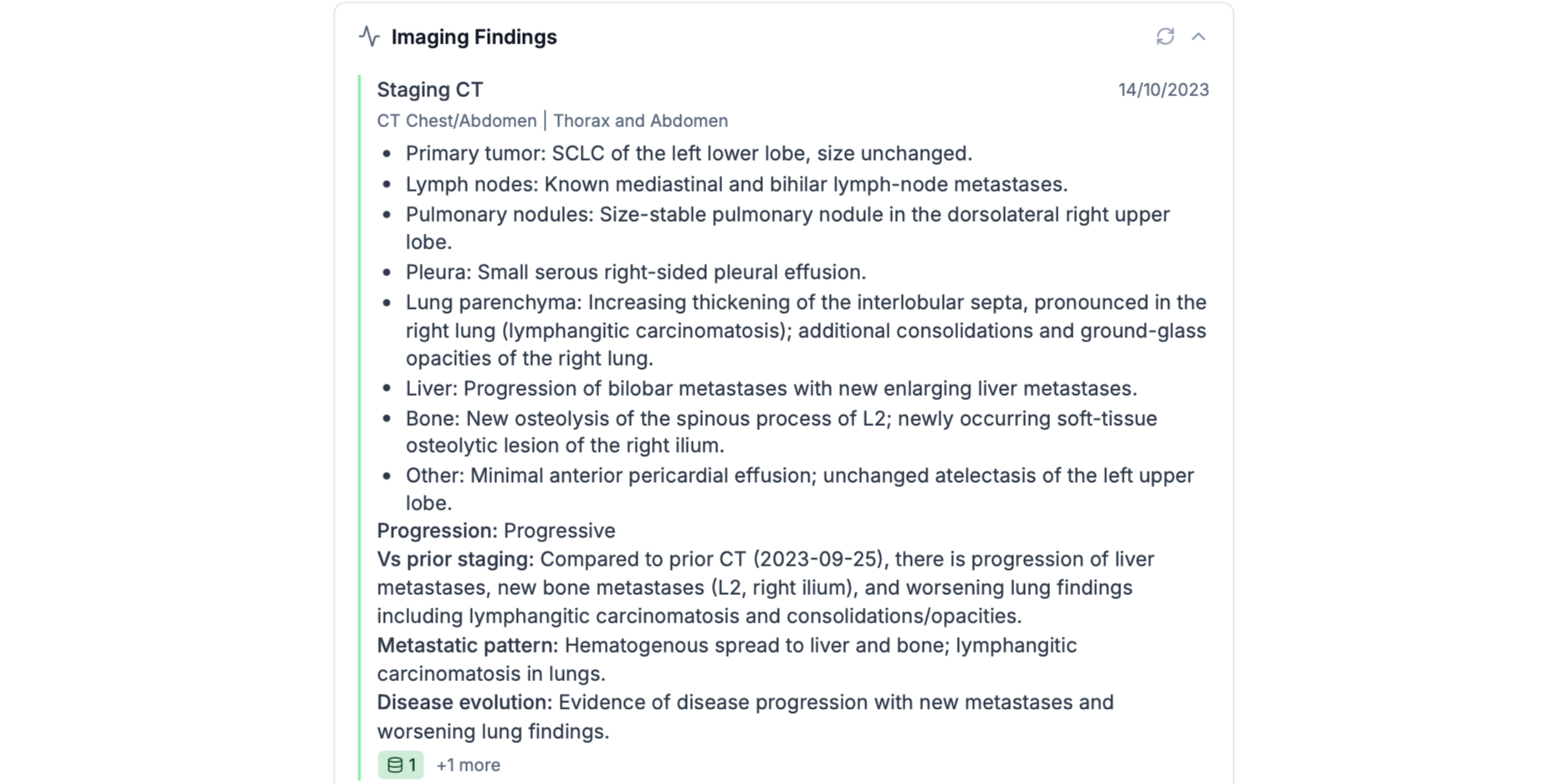
Disease-course-aware timeline. TURBO retrieves structured and unstructured data from a FHIR server and reconstructs a chronological timeline of the patient's medical history, including LLM-derived key insights per event. In addition, MedGemma derives treatment responses over time from imaging reports.

Structured cross-document synthesis. MedGemma integrates radiology reports, pathology, therapies, and comorbidities into a coherent case summary. Statements are linked to their source resources, enabling rapid comprehension with traceability. Rather than producing a generic summary, the model synthesizes information across longitudinal records in alignment with tumor board decision-making needs.
Agentic interaction with tool access. TURBO's tool-enabled agent can retrieve additional FHIR resources and query guideline content from a vector database. This allows clinicians to ask targeted questions about the patient in a chat interface. In addition, the agent identifies missing information for the tumor board, such as the patient's wishes regarding further therapies, and requests that the clinician complete the case summary.
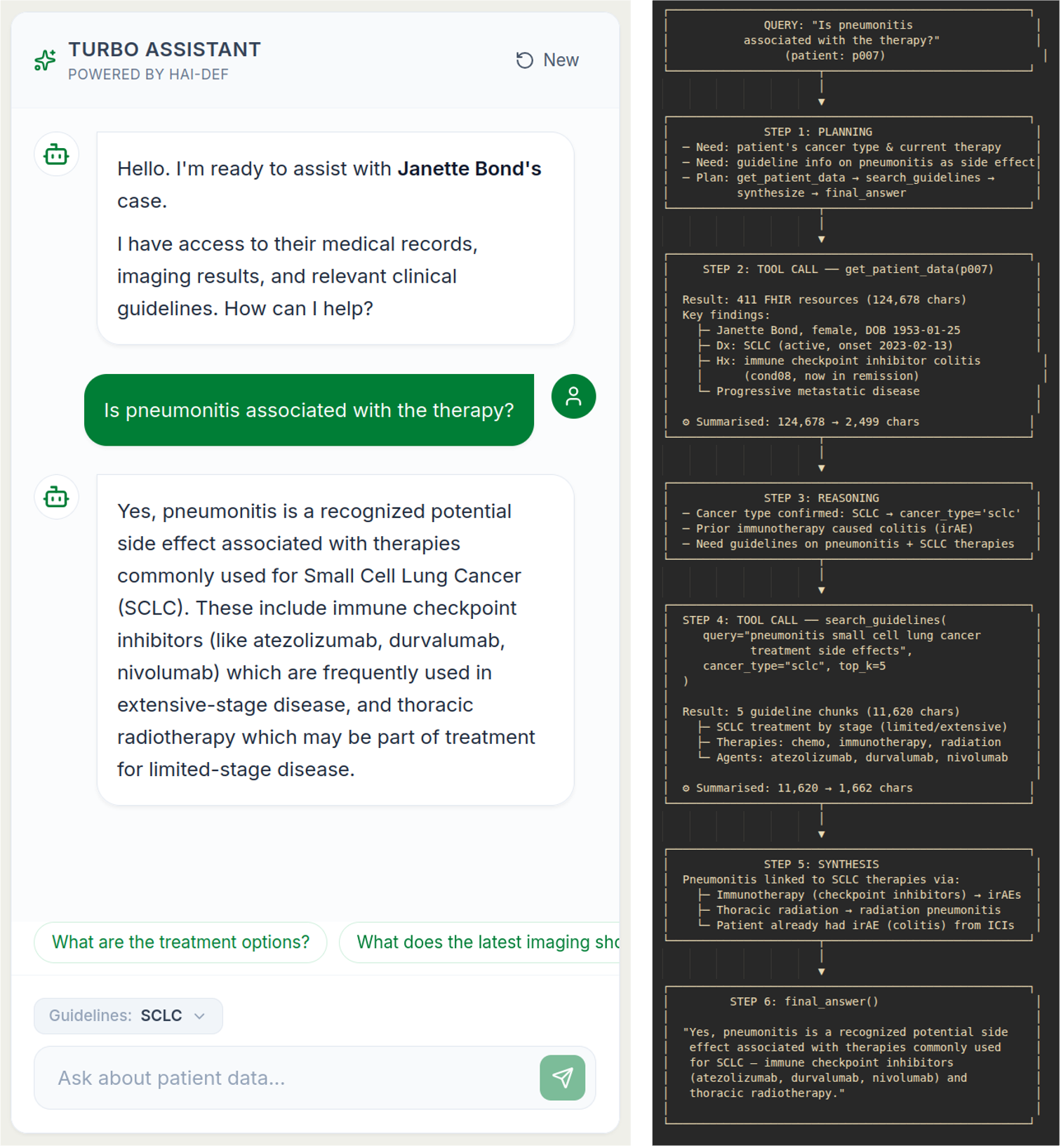
For all components, MedGemma is essential because the task requires domain-specific language understanding and long-context reasoning, as well as secure on-premise deployment. Its medical pre-training and open-weight architecture enable clinically grounded reasoning within hospital infrastructures. This makes TURBO a deployable workflow solution rather than a standalone AI demonstration.
TURBO is implemented as a modular, self-hostable architecture.
System architecture:
Frontend: React + Tailwind dashboard for patient selection, structured summary display, timeline visualization, and interactive chat.
Backend: FastAPI service orchestrating FHIR retrieval and normalization, prompt construction, and agent tool-calling.
- Clinical data repository: HAPI FHIR R4 server providing structured clinical resources (diagnoses, procedures, observations, reports).
- Knowledge database: Milvus vector database storing embedded medical guideline documents.
- LLM layer: On-premise MedGemma instance deployed within the hospital network and accessed via an OpenAI-compatible API endpoint. If such an endpoint is used, all inference calls are performed locally without external data transfer.
- Agent framework: smolagents with MCP tools enabling controlled access to FHIR queries and guideline retrieval.
Deployment: Docker Compose for a reproducible setup across the frontend and backend.
Workflow implementation:
- A physician selects a patient in the dashboard.
- The backend retrieves and normalizes relevant FHIR resources.
- The backend calls the local MedGemma API to generate:
- a longitudinal timeline with key insights per event and information on treatment response
- a structured case summary,
- The agent supports follow-up queries via chat interface, performs targeted FHIR retrieval, and links statements to their source resource.
All model outputs are explicitly source-referenced to ensure transparency and verifiability.
Feasibility and next steps The stack is fully self-hostable and aligned with hospital data governance constraints. The current MVP runs on a preconfigured FHIR server with synthetic data in Docker Compose. Next steps include prospective measurement of time savings against manual preparation, quantitative evaluation of summary completeness and citation accuracy, systematic assessment of hallucination rates across case complexity levels, and reliability hardening for routine clinical use.
[1] Schreyer AG et al. Interdisziplinäre Tumorkonferenzen in der radiologischen Routine. Der Radiologe 2020;60:737–746.
[2] Lajmi N et al. Simulation-Based Evaluation of a Large Language Model–Enabled Clinical Decision Support Platform in Oncology. JCO Clin Cancer Inform 2026;10:e2500244.
[3] Roche Diagnostics. AI in oncology: Improving tumor board workflows. Healthcare Transformers, November 2025. https://diagnostics.roche.com/global/en/healthcare-transformers/article/ai-in-oncology-tumor-boards.html
[4] Bray F, Laversanne M, Sung H, et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2024;74(3):229–263.
[5] El Saghir NS et al. Global Practice and Efficiency of Multidisciplinary Tumor Boards: Results of an ASCO International Survey. JCO Global Oncol 2015;1:57–64.
