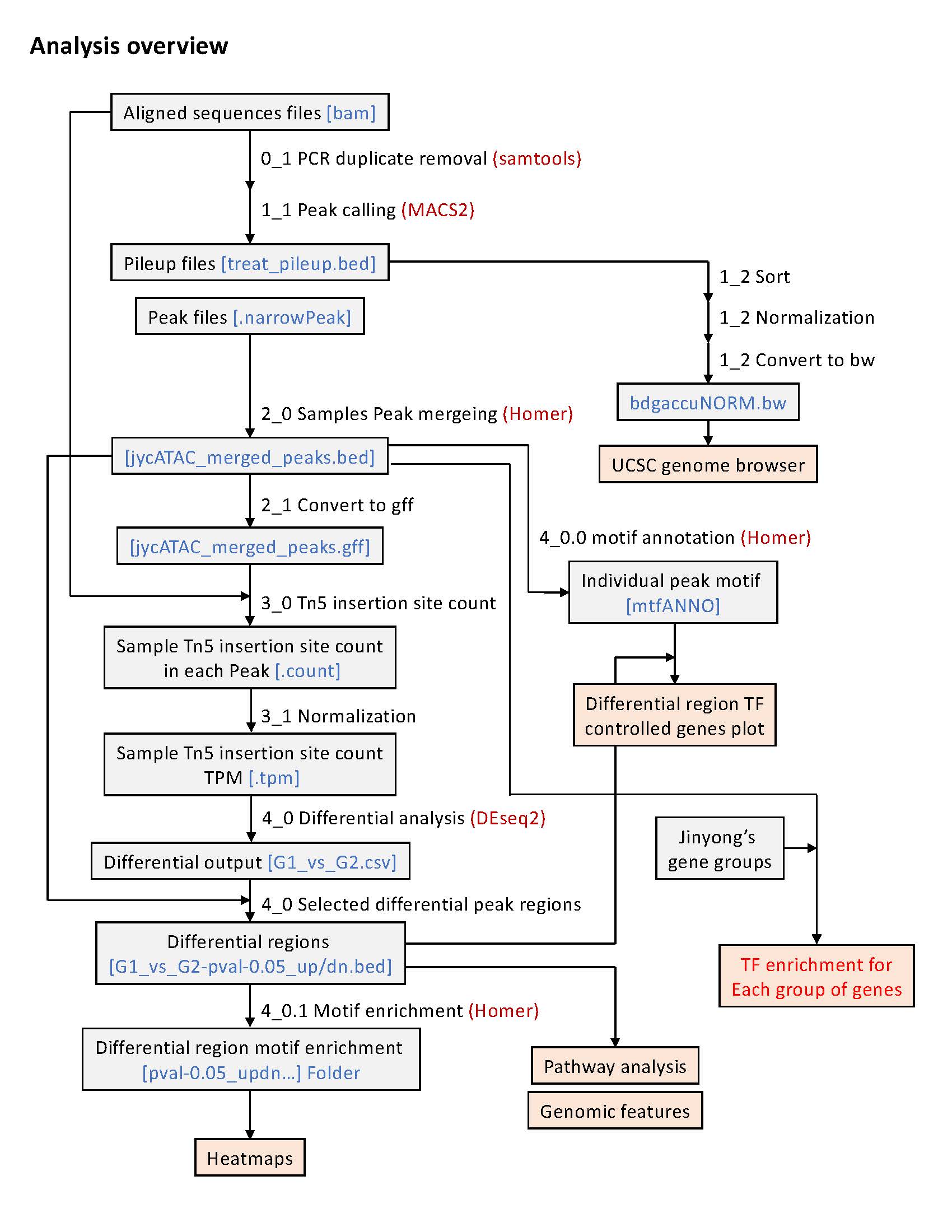
- All ATAC-seq bdg files (then converted to bw) normalized by scaling_factor calculated by:
accumulation=($(awk '{sum+=(($3-$2)*$4)} END {print sum}' $input_bdg_file))
scaling_factor= 1e+09 / $accumulation
Folder description:
[Folder] 9_Figures_plottingCodes/0_motifHeatMap
0_only_homer_known: motif enrichment heatmaps with only homer known motif
1_combined_motifs: motif enrichment heatmaps created with combined homer known motif + JYC motif
Motif enrichment percentage in significant differential regions (DEseq2 pval < 0.05)
Naming:
Condition1_vs_Condition2_up: Regions that are more accessible in condition1 than condition2
all_sig_motifs_c25: Motifs that are selected based on > 25% enrichment in any group
byfamily: Spreadsheet for motif enrichment ranked by TF family
Motif enrichment in significant differential regions of Comparison1 v.s. significant differential regions of Comparison2
Naming:
[Filename]vs−WT_Tfh_vs_Naive_dn_XXXX.pdf: All the differential region motif enrichments conditions are compared to enrichment in WT_Tfh_vs_Naive_dn regions (WT Tfh less accessible than Naiv)
Folder description:
[Folder] 6_GeneGroupMotifAnalysis
byGroup: motif analysis of peaks annotated to genes in each group
byGroup_open: motif analysis of peaks annotated to genes in each group & more accessible in regions that are more accessible in Tfh/Th1 than Naive
- Main resullts folder: 6_GeneGroupMotifAnalysis/byGroup_open/vsNAVopen_cb_mtfs_motif-finding
- System: Ubuntu, nonGUI, x86_64 Cyverse Atmosphere Server
- Pre-requisites: